Genomics England uses MongoDB to Power the Data Science Behind the 100,000 Genomes Project
MongoDB Enterprise Advanced Scales to Handle Hundreds of Terabytes of Complex Genetic Data to Power Life Saving and Groundbreaking Discoveries
MongoDB Enterprise Advanced Scales to Handle Hundreds of Terabytes of Complex Genetic Data to Power Life Saving and Groundbreaking Discoveries
London, UK – April 10, 2018 – Genomics England is using MongoDB (Nasdaq: MDB), the leading modern, general purpose data platform, to power the data science that makes the 100,000 Genomes Project possible and reduce the processing time for complex queries from hours to milliseconds, enabling scientists to discover new insights, faster.
Genomics England, a company owned by the UK government’s Department of Health and Social Care, runs the 100,000 Genomes Project. This flagship project is sequencing 100,000 whole genomes from patients with rare diseases and their families as well as patients with common cancers. Working with the UK’s National Health Service, the aim is to harness the power of whole genome sequencing technology to transform the way people are cared for. Patients may be offered a diagnosis where there wasn’t one before. In time, there is the potential of new and more effective personalized treatments for patients.
Patients’ genomic data is combined with their clinical data to enable interpretation and analysis. Approved researchers, clinicians and industry partners can also access the Project’s dataset – studying how best to use genomics in healthcare and how best to interpret the data to help patients.
The project is sequencing on average 1,000 genomes per week, producing about 10 terabytes of data per day. To manage this complex, massive and sensitive data set, Genomics England uses MongoDB Enterprise Advanced as an important component of its computing platform.
“Managing clinical and genomic data at this scale and complexity has presented interesting challenges,” said Augusto Rendon, Director of Bioinformatics at Genomics England.
“That’s why adopting MongoDB has been vital to getting the 100,000 Genomes Project off the ground. It has provided us with great flexibility to store and analyze these complex data sets together. This will ultimately help us to realize the benefits of the Project - delivering better diagnostic approaches for patients and new discoveries for the research community."
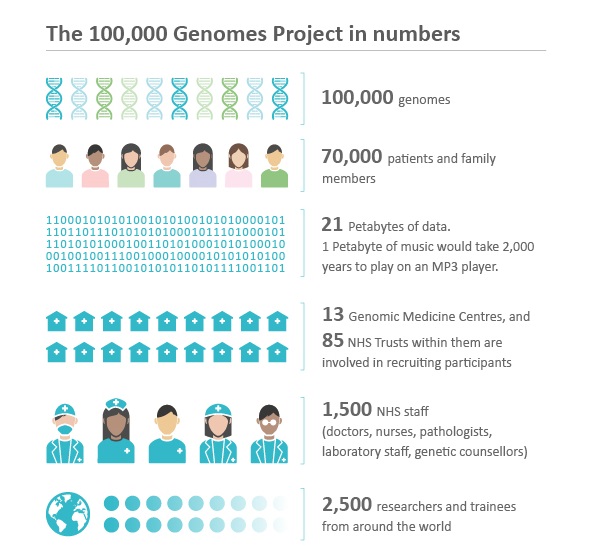
Credit: Genomics England
Why MongoDB
The project’s data platforms were built from the ground up on MongoDB. The document database was chosen for three core reasons. The first is the database’s native ability to handle a wide variety of data types, even those data structures that weren’t considered at the beginning of the project. This makes it simple for developers and data scientists to rapidly evolve data models and develop software solutions. The second reason was performance at scale. It was clear there would be a massive and constantly growing data set that would need to seamlessly scale across underlying compute resources. Not only would the data set be big and complex, but researchers need to easily explore the data and not have long waits for simple queries.
The third key driver for Genomics England to select MongoDB was its operational maturity with features such as end-to-end encryption, fine-grained access control to data and operational tooling. The 100,000 Genomes Project dataset is sensitive: as well as the full genetic makeup of a patient, it also includes their clinical features and lifetime health data. Instead, de-identified data is analyzed within a secure, monitored environment. This makes encryption and data security vital. Varying levels of complex permissions are required so that only those with the appropriate credentials can access the most sensitive data sets.
MongoDB Enterprise Advanced satisfied these requirements and has been providing Genomics England with data flexibility, performance at scale and security since the project started in 2013.

Credit: Genomics England
Ignacio Medina, Head of Computational Biology Lab HPC Service, University of Cambridge, and Head of Bioinformatics Databases at Genomics England has been building many of the applications that sit on top of MongoDB. He said, “MongoDB is performing beautifully for us. From the beginning of the project it’s been fantastic for our developers to build and iterate quickly. Now that the 100,000 Genomes Project is running at scale, MongoDB is also helping us extend that great experience on to the scientists and clinicians who access the data, making it easier and faster for them to find critical insights in the data.”
Two of the most important projects are Cellbase and OpenCGA (Computational Genomics Analysis). Cellbase is a data warehouse and open API that stores reference genomic data from public resources such as Ensembl, Clinvar, and Uniprot. The genomic information itself includes the specific annotations other researchers have made on cell mutations. The code for Cellbase is available on Github. By relying on MongoDB, Cellbase can typically run sophisticated queries in an average of 40 milliseconds or less, and complex aggregations in less than 1 second. This is down from six hours using previous filesystem-based querying and storage. Importantly, it can annotate about 20,000 variants per second, making it compatible with whole genome sequencing data throughput requirements, while also returning a rich set of annotations that helps scientists better understand the data.
OpenCGA aims to provide researchers and clinicians with a high-performance solution for genomic big data processing and analysis. The OpenCGA platform also includes much more detailed information on genomic material including private metadata and more specific background information on specific sequenced genomes. This means OpenCGA has the ability to process incredibly complex queries based on a huge variety of variables, from a single mutation right through to analyzing common mutations within a specific geographic region.
By using MongoDB, OpenCGA enables researchers to query data in any way they want. This is done by use of MongoDB’s secondary indexes – from compound indexes to query data across related attributes, text search facets to efficiently navigate and explore data sets, and sparse indexes to access highly variable data structures. Each collection can have 20 more secondary indexes to service multiple query patterns, including complex, ad-hoc queries.

Credit: Genomics England
The project has now reached its halfway point, with over 50,000 genomes sequenced and patients already benefiting from new diagnoses and opportunities to take part in clinical trials. By the end of 2018 the 100,000 Genomes project will be complete, at which point there will be more than 20 petabytes of data stored on the project’s infrastructure. The project will go on to provide the infrastructure for a National Genomics Service being put in place by NHS England.
Dev Ittycheria, President and CEO, MongoDB, concluded, “The 100,000 Genomes Project hits home for me in a very personal way as I recently lost my mother to cancer. I am extremely grateful that so many brilliant people are dedicating their time and energy to this important project. We are honored that MongoDB is playing an essential role as the underlying data platform to produce data science that is likely to change the lives of millions of people, including someone we may personally know, for the better. This is the kind of project that inspires us to do our best work every day.”
About Genomics England
Genomics England is a company owned by the Department of Health and Social Care, and was set up to deliver the 100,000 Genomes Project. This flagship project will sequence 100,000 whole genomes from NHS patients and their families.
Genomics England has four main aims:
- to bring benefit to patients
- to create an ethical and transparent programme based on consent
- to enable new scientific discovery and medical insights
- to kickstart the development of a UK genomics industry
The project is focusing on patients with rare diseases, and their families, as well as patients with common cancers. For more information visit www.genomicsengland.co.uk.
About MongoDB
MongoDB is the leading modern, general purpose database platform, designed to unleash the power of software and data for developers and the applications they build. Headquartered in New York, MongoDB has more than 5,700 customers in over 90 countries. The MongoDB database platform has been downloaded over 35 million times and there have been more than 850,000 MongoDB University registrations.